A Bioinformatics Approach for the Phenotype Prediction of Nonsynonymous Single Nucleotide Polymorphisms in Human Cytochromes P450 | Drug Metabolism & Disposition

Inferring the molecular and phenotypic impact of amino acid variants with MutPred2 | Nature Communications

Filtrating steps to identify variant associated with the phenotype.... | Download Scientific Diagram

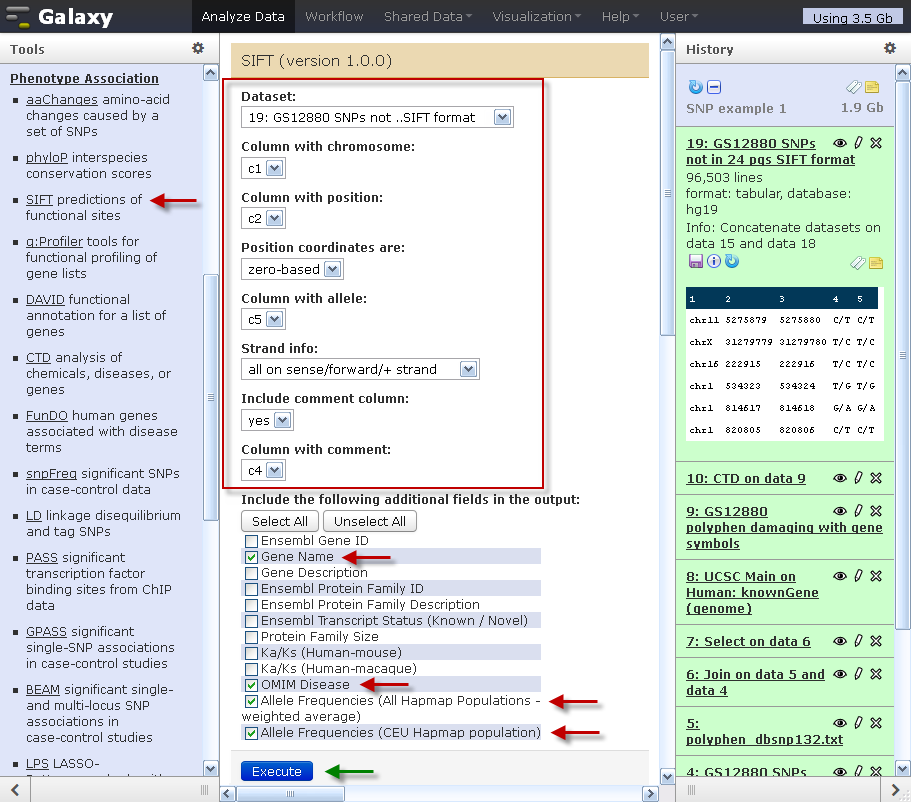
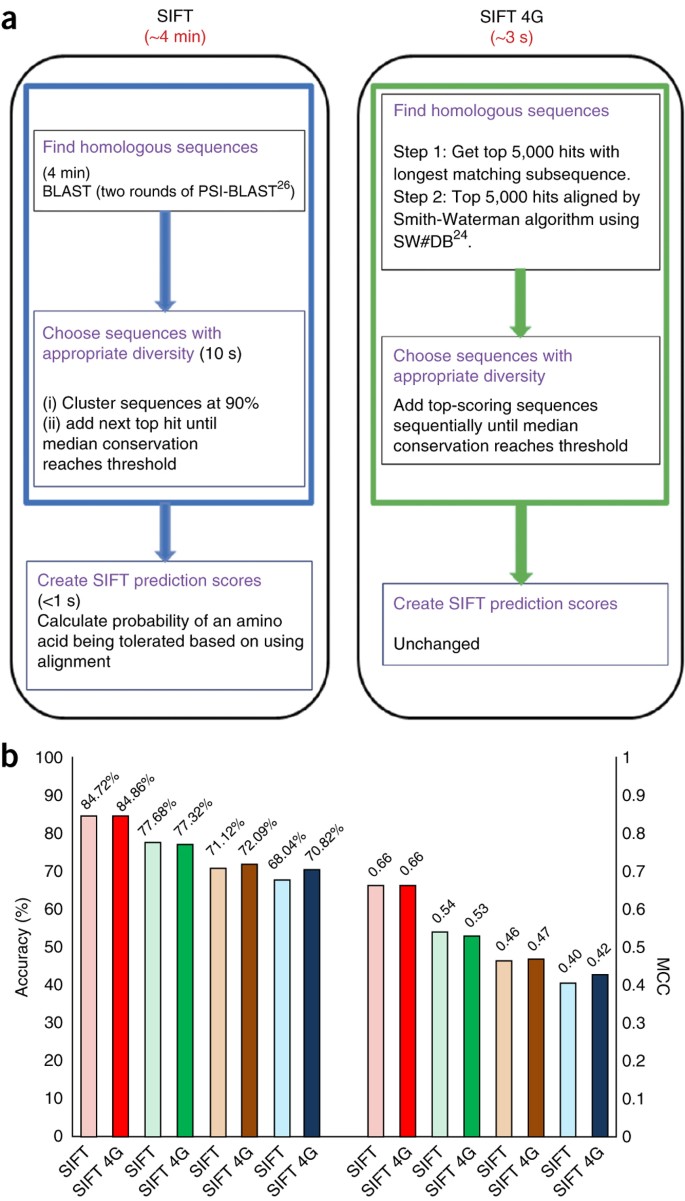
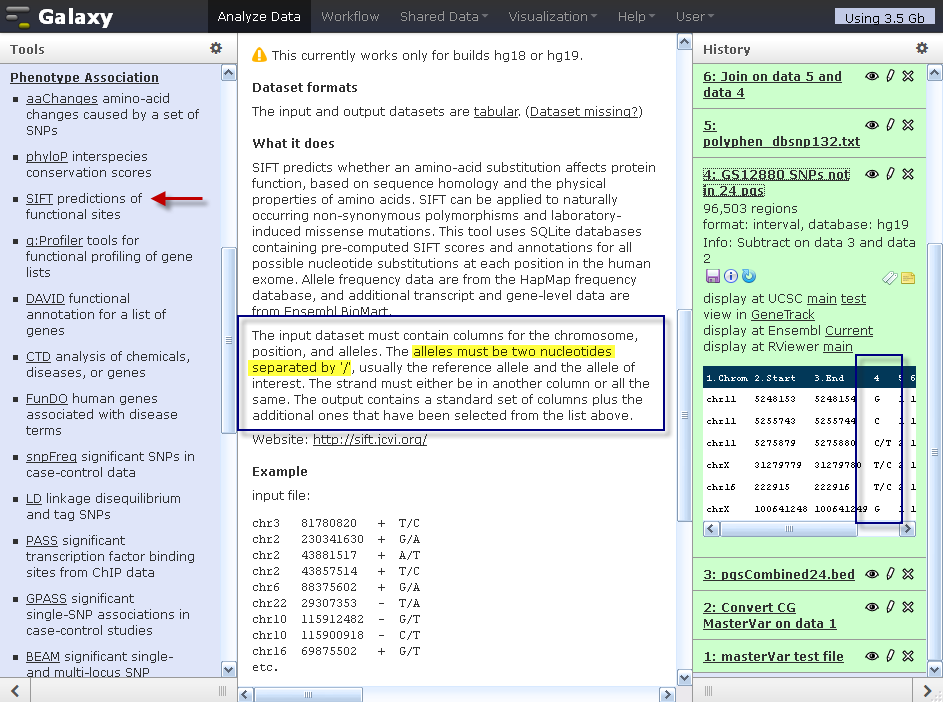
Using SIFT and PolyPhen to Predict Loss-of-Function and Gain-of-Function Mutations | Genetic Testing and Molecular Biomarkers
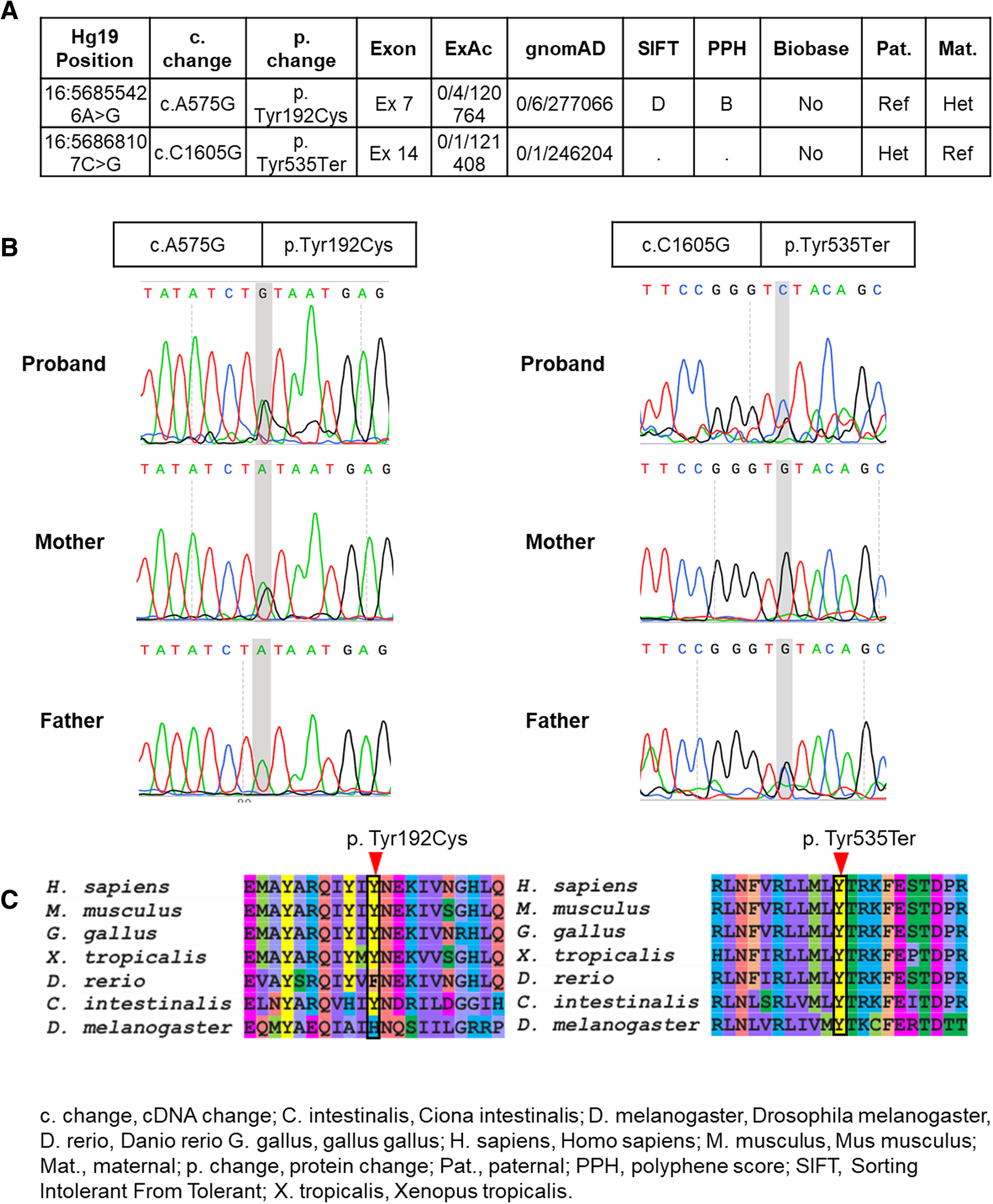
Identification of novel mutations and phenotype in the steroid resistant nephrotic syndrome gene NUP93: a case report | BMC Nephrology | Full Text

Bridging Genomics to Phenomics at Atomic Resolution through Variation Spatial Profiling - ScienceDirect
Population-Based Resequencing of APOA1 in 10,330 Individuals: Spectrum of Genetic Variation, Phenotype, and Comparison with Extreme Phenotype Approach | PLOS Genetics
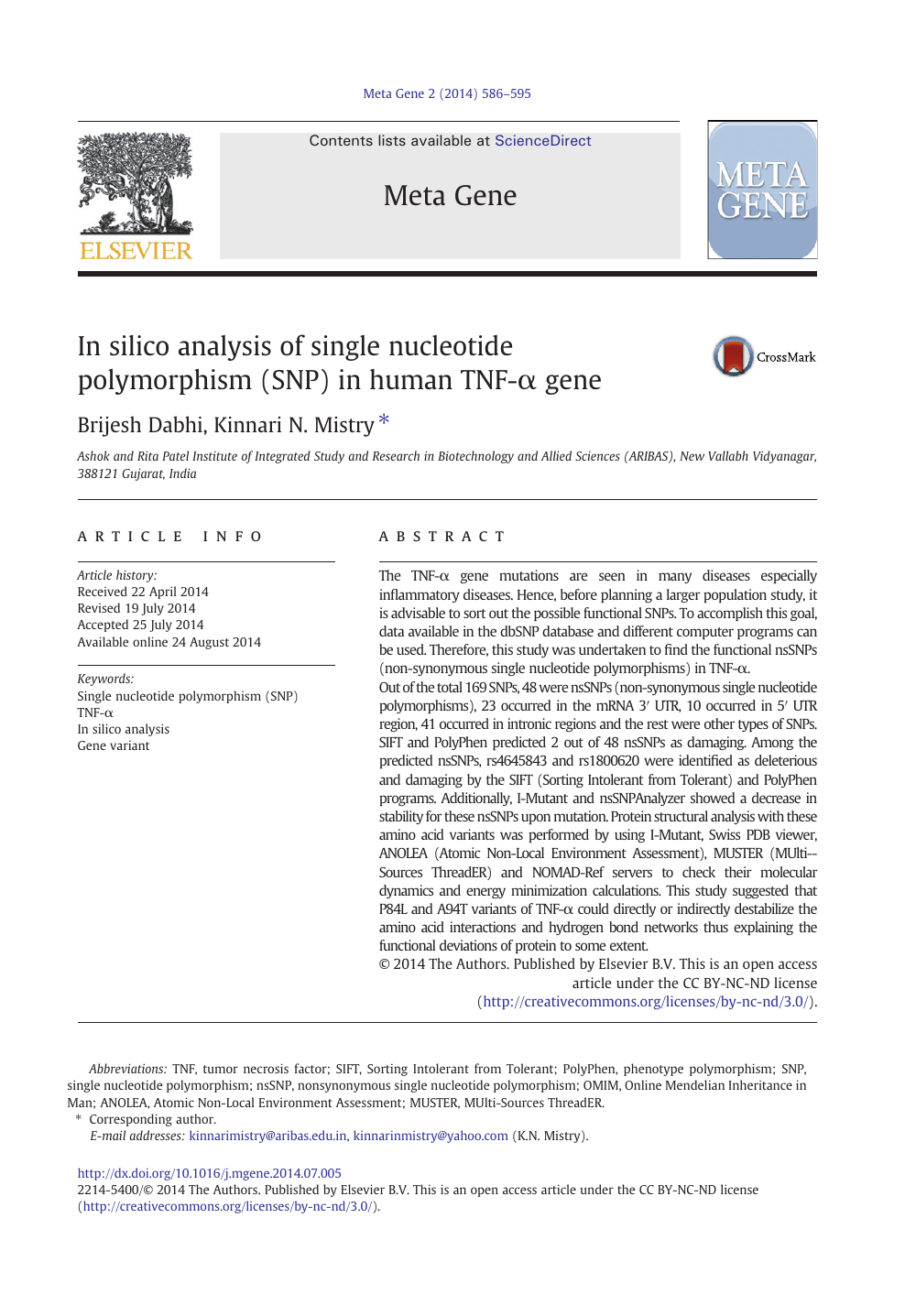

In silico analysis of single nucleotide polymorphism (SNP) in human TNF-α gene – topic of research paper in Biological sciences. Download scholarly article PDF and read for free on CyberLeninka open science

In silico analyses of missense mutations in coagulation factor VIII: Identification of severity determinants of haemophilia A - University Of Calcutta
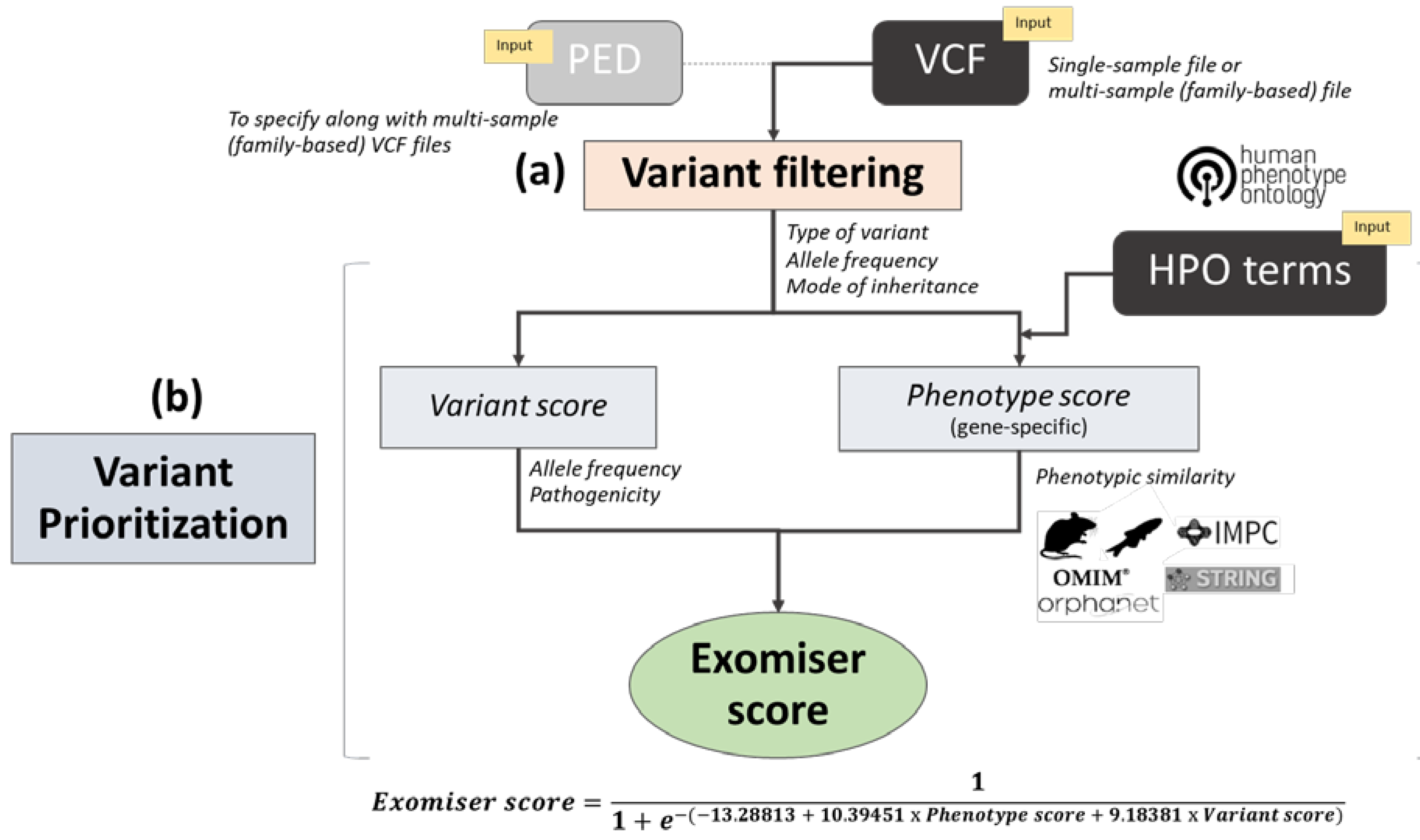
Genes | Free Full-Text | An Improved Phenotype-Driven Tool for Rare Mendelian Variant Prioritization: Benchmarking Exomiser on Real Patient Whole-Exome Data

Expansion of the mutation spectrum and phenotype of USP7-related neurodevelopmental disorder | Semantic Scholar